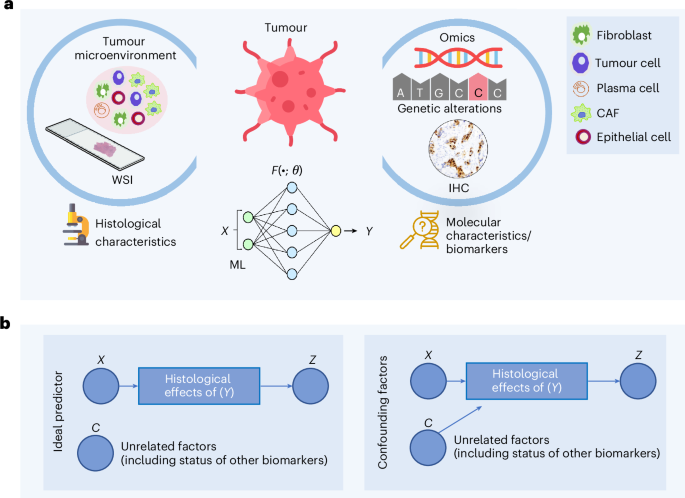
Deep learning models that infer clinically relevant biomarker status from tissue images are being explored as rapid and low-cost alternatives to molecular testing. Here we show, through statistical analysis across multiple cancer types, datasets and modelling approaches, that the datasets used to train these models contain strong dependencies between biomarkers and clinicopathological features, which prevent models from isolating the effect of a single biomarker and lead them to learn confounded signals. Consequently, their prediction accuracy varies substantially with the status of codependent biomarkers and clinicopathological variables, and for several biomarkers, the gain over what a pathologist can already infer from routine histopathological features, such as grade, remains modest. These findings indicate that current approaches are not yet suitable as substitutes for molecular testing but can support triage or complementary decision-making with caution. Unconfounded biomarker prediction will require models that learn causal rather than correlational relationships between biomarkers and tissue morphology. This study shows that biomarker prediction accuracy from whole-slide images can reflect confounding by codependent biomarkers and correlated variables, suggesting that models may not learn the intended biology and may face challenges in clinical translation. Data and study design We analysed the limitations of existing ML approaches for predicting molecular biomarkers (for example, mutations, genomic instability indicators and protein expression) from H&E stained WSIs. A high-level concept diagram of the... [44100 chars]