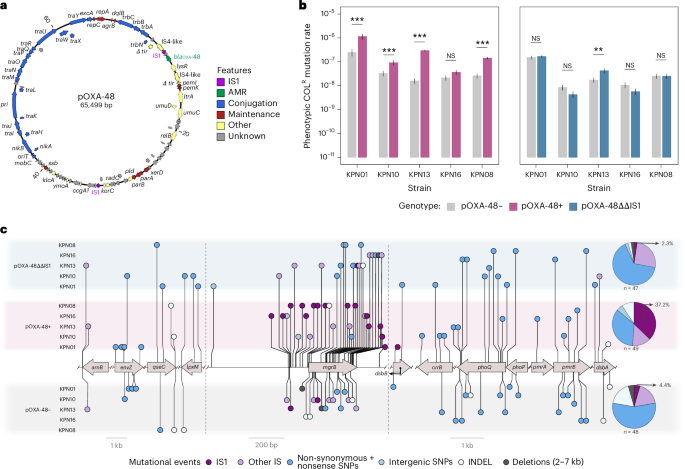
Plasmids are mobile genetic elements that can rapidly spread across bacterial populations, promoting the dissemination of antimicrobial resistance (AMR) genes in bacteria. They are enriched in insertion sequences (IS), which are small transposable elements able to translocate between genetic locations. Here we combined experimental, bioinformatic and computational approaches to investigate the role of plasmids promoting AMR through IS-mediated gene inactivation. We find that plasmid pOXA-48, which encodes two IS1 elements, increases the rate of resistance acquisition to multiple antibiotics in clinical strains of Klebsiella pneumoniae through IS1-mediated gene disruption. Screening of genome databases confirmed that the inactivation of genes through plasmid-encoded IS elements is an extended mechanism of AMR evolution. Both our experiments and computational model revealed that conjugative plasmids can promote this route of AMR acquisition while invading complex bacterial communities. Overall, we show that conjugative plasmids contribute to AMR not only through the dissemination of resistance genes, but also through IS-mediated gene inactivation. Inactivation of chromosomal genes through plasmid-encoded IS elements is an extended mechanism of antimicrobial resistance evolution in bacteria. All mutations detected across all conditions and strains. Mutations accumulated in the absence of pOXA-48 are represented on the lower part of the panel (grey background). Mutations accumulated in the presence of pOXA-48 (purple background) or pOXA-4... [6373 chars]