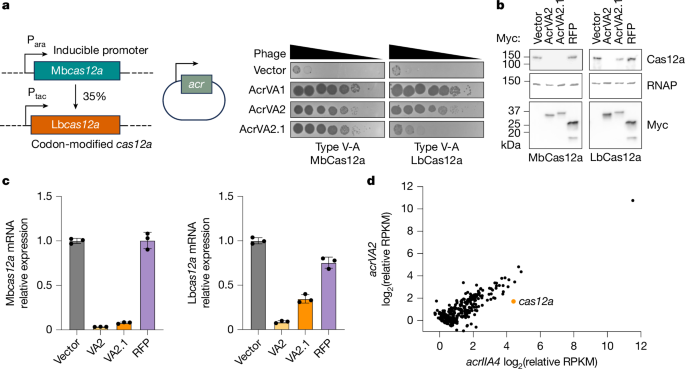
Bacteria encode diverse defence systems, including CRISPR–Cas, to recognize and cleave the DNA of bacteriophages (phages) and other mobile genetic elements1. In response, phages encode anti-CRISPR (Acr) proteins that inhibit CRISPR–Cas activity by blocking DNA binding or cleavage2. Here we report an unexpected mechanism by which the anti-CRISPR AcrVA2 inhibits Cas12a biogenesis. AcrVA2 binds conserved and functionally important amino acid residues near the Cas12a N-terminus and triggers selective degradation of cas12a mRNA as it is translated. Additionally, conserved residues in the AcrVA2 C-terminal domain enable co-sedimentation with ribosomes and polysomes, which is required to achieve targeted co-translational mRNA degradation. The AcrVA2 C-terminal domain is broadly conserved in homologs encoded by diverse mobile genetic elements, typically in hosts that lack cas12a, suggesting that these homologues may recognize and downregulate alternative substrates in other bacteria. These findings reveal a novel mechanism for molecular conflict and gene regulation in bacteria. The anti-CRISPR protein AcrVA2 specifically interrupts Cas12a biogenesis by triggering co-translational mRNA degradation. Extended Data Fig. 1 AcrVA2 inhibits and downregulates diverse variants of cas12a. (a) Phage plaque assay using ten-fold serial dilution of phage to assess Cas12a inhibition by AcrVA2 (or homolog AcrVA2.1) compared to controls. 33362, 237, and 58069 ... [8487 chars]